Ah, let me take you on a nostalgic journey back to my days as an academic researcher. Among the studies I conducted, there’s one that still fills my heart with hope. It’s about these funky noncovalent interactions in protein structures called anion-π. In a previous story, I’ve shared my recollection on personal experiences, scientific collaborations, and the stubbornness that went into that study. Since its publication in 2011, I’ve been keeping an eye out for other works that reference it, and just recently, I’ve stumbled upon this awesome paper by Adhav & Saikrishnan. They did a great job at categorizing what they call “unconventional” noncovalent interactions in proteins. Low barrier hydrogen bonds, σ-hole, anion-π… let’s dive into all these chemical oddballs and figure out how they contribute to the structure stability and functions of proteins.

Noncovalent Interactions in proteins
You remember the unsung heroes of the molecular world we talked about in From Atoms To Words, right? Yes, proteins. These fantastic catalytic machines keep life rolling by silently carrying out vital tasks. And guess what? The secret to how a protein works lies in its 3D structure.
Think of the 3D structure as the protein’s blueprint. It calls the shots on what the protein can do and how it gets things done. Any deviation or disruption in that structure can throw cellular activities off balance, leading to hellish physiological disorders and diseases. It’s like a chaotically evil domino effect.
That’s why it’s absolutely crucial for proteins to adopt and maintain their functionally relevant structures. It’s about finding that sweet spot where everything falls into place flawlessly. And you wanna know who’s shaping those 3D wonders?
Noncovalent interactions, my friend.
But hold on, you say: What the heck are noncovalent interactions?
Countless volumes have been written about it. But let’s try to answer in a few lines.
Imagine two atoms getting cozy and deciding to share their electrons with each other. It’s like a trust fall, where the electrons happily bounce in between atoms. They’re in it together, folks. That’s what makes a covalent bond—the electron-sharing fiesta.
Noncovalent interactions are different. These interactions, my curious friend, take shape either by a complete exchange of electrons between atoms (ionic bonding like in our kitchen salt for example) or by not exchanging any electrons at all. Within this latter case, we encounter conventional examples, such as hydrogen bonds and van der Waals interactions.
More on From Atoms To Words:
▸ From Earth to the Cosmos: How Hydrogen Bonds Shape Life
▸ When Will RNA Structure Prediction Get Its AlphaFold Breakthrough?
▸ All-Atom Molecular Dynamics of SARS-CoV-2: The Computational Microscope’s View of 305 Million Atoms
In proteins, covalent bonds (known as peptide bonds) play their part in linking amino acids together, but it’s the noncovalent interactions that add the enchantment of three-dimensionality. They fold those polypeptide chains into precise, well-defined structures, giving proteins their unique shapes. It’s like witnessing an intricate origami masterpiece, meticulously guided by these interactions.
But noncovalent interactions aren’t just about structure; they influence protein function as well. They impact how proteins bind to ligands, form complexes with other proteins, nucleic acids, lipids, membranes, and carbohydrates. They’re the mastermind behind molecular recognition, enabling proteins to identify and interact with specific partners. And let’s not forget their vital role in enzymatic reaction mechanisms and kinetics.
The reason noncovalent interactions are such key players in the protein game is because of their incredible ability to break and form with ease. Unlike strong and rigid covalent bonds, noncovalent interactions are flexible as their strength varies, roughly from 0.5 to 50 kcal/mol, depending on the atoms or molecules involved and the environment they’re in.
The bottom line is: if we truly want to understand proteins inside out, we’ve got to dive headfirst into the world of noncovalent interactions.
So, my friend, today we embark on a journey inspired by the solid article by Adhav & Saikrishnan. Let’s plunge into the realm of these funky, unconventional noncovalent interactions.

7 Unconventional noncovalent interactions in proteins
1. Low barrier hydrogen bonds
These hydrogen bonds – they’re a bit special. You see, they form when the acidity of the hydrogen bond donor and acceptor atoms are on par, meaning they can both let go of a hydrogen proton. As a result, the hydrogen atom sits right in the middle, keeping’ things symmetrical. Energetically this produces a low energy barrier between two minima that are equally deep. Picture it as a gentle slope connecting two valleys. And guess what? The hydrogen can easily hop from one valley to the other without breaking a sweat.
Compared to your run-of-the-mill regular hydrogen bonds, these unconventional interactions in proteins are usually stronger (12-24 kcal/mol). This extra strength can have a profound impact on the kinetics of various biological processes, such as protein folding, molecular recognition, and ligand binding.
2. C5 hydrogen bond
This kind of interaction forms between an oxygen of a carbonyl (C=O) group and the nitrogen of the amide group, all within the same amino acid. That’s why we call it the intraresidue C5 hydrogen bond.
Fancy, right?
This C5 hydrogen bond is no stranger to the protein party. In fact, it’s pretty common, as it shows up in 94% of proteins structures and especially in the protein substructures known as β-sheets, where it’s present in about 60% of cases. Its role? Contributing to the stability of the structures it forms in.
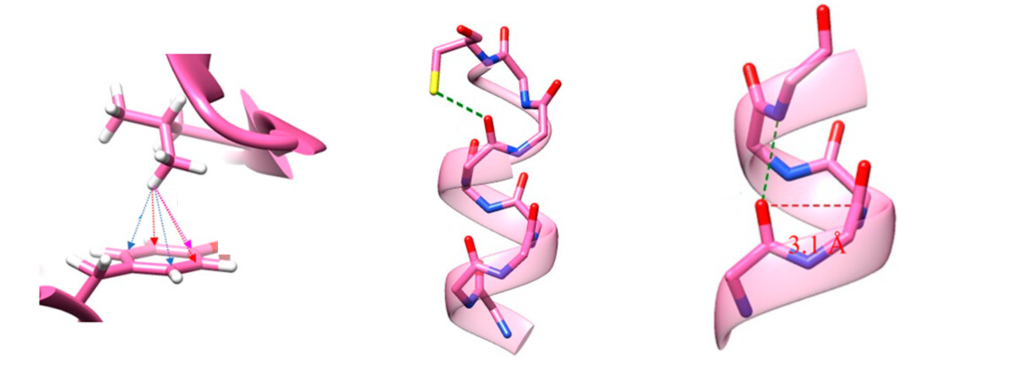
3. C-H…π interactions
This is a type of noncovalent interaction where a C-H group steps up as a hydrogen bond donor, and those sweet π electrons from an aromatic ring (think benzene-like molecules) play the role of the acceptor. It’s like a cool geometric cousin to other hydrogen bonds, and its strength, according to quantum chemistry calculations, ranges between 1.5 and 2.5 kcal/mol.
Protein aromatic rings with their π electrons absolutely adore this interaction as involves around three-quarters of tryptophans, half of phenylalanines and tyrosines, and a quarter of histidines. Although relatively weak, our humble C-H…π interactions are believed to favor protein folding and to contribute to the overall stability of our untiring catalytic machines.
But hold on tight, because there’s more to this story. These interactions have been spotted frequently in protein-DNA complexes, suggesting they have a role in the intricate dance of protein-DNA binding and recognition.
More on From Atoms To Words:
▸ AI in Drug Discovery: Chasing Dreams, Facing Realities
▸ Is Machine Learning Going to Replace Computational Chemists?
▸ The Evolution of Quantum Chemistry: From Pencil and Paper to Quantum Computing
4. Sulfur-mediated hydrogen bonds
As the name implies, these hydrogen-bond-like interactions involve sulfur-containing amino acids like cysteine and methionine. What’s interesting about it is that sulfur, with its relatively low electronegativity, doesn’t pack as much punch compared to other elements. So, these bonds tend to be on the weaker side, with energies hovering around 5 kcal/mol.
But don’t let their weakness fool you, because these sulfur-mediated hydrogen bonds can still have a significant impact on protein structure and function. For example, this modest connection is responsible for stabilizing α-helices, those cool spiral-like structures we see in proteins. Or again, messing with one of them in iron-sulfur proteins can actually affect the protein’s activity itself. It’s like taking a spicy ingredient out of a recipe.
5. n → π* interactions
So, you’ve got a carbonyl group (C=O), where the oxygen carries a partial negative charge, while the carbon is rocking a partial positive charge. Now, when you’ve got two of these carbonyl groups hanging out together (C=O···C=O), they start to interact.
This interaction gets its funky name because there’s a charge transfer going on. The electron density from the oxygen lone pair (n) hops over to the antibonding orbital (π*) of the carbonyl group. This creates a pretty weak interaction, clocking in at less than 1 kcal/mol. Doesn’t sound like much, right? But considering how often this interaction occurs, it actually adds up and contributes to the overall stability of proteins, especially those beautiful α-helices.
6. σ-hole interactions
Ever heard of a σ-hole? I bet you’re curious. Well, let me fill you in. It’s the outer lobe of a half-filled p orbital, which plays a key role in the formation of covalent bonds. However, when this orbital finds itself short of electrons, something interesting happens. It creates a zone buzzing with positive electrostatic potential. And you know what? This potential can’t resist a little rendezvous with negative molecular sites, sparking an attractive (yet noncovalent) interaction. And that’s what we refer to as σ-hole bonding.
Now, depending on the characteristics of the players involved, we’ve got ourselves a variety of σ-hole bonding flavors. Let’s start with halogen bonding, where our trusty companions chlorine, bromine, and iodine atoms take the stage. Then there’s chalcogen bonding interaction, starring sulfur, selenium, and tellurium. And we mustn’t overlook tetrel bonding, featuring the likes of carbon, silicon, germanium, tin, or phosphorus.
These distinct types of σ-hole bonding bring their own unique vibes to the realm of protein stability, adding a touch of complexity to the intricate network of molecular interactions.

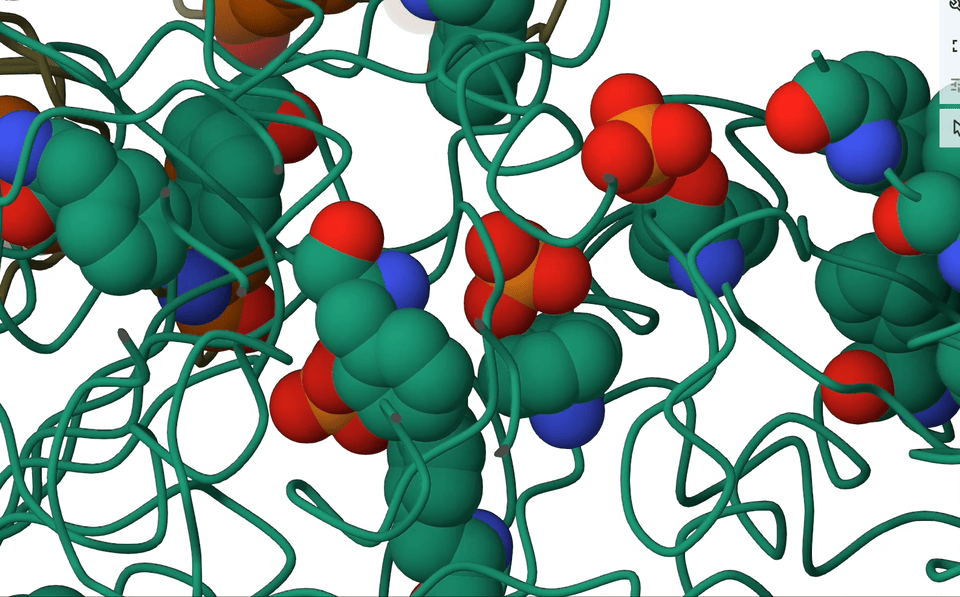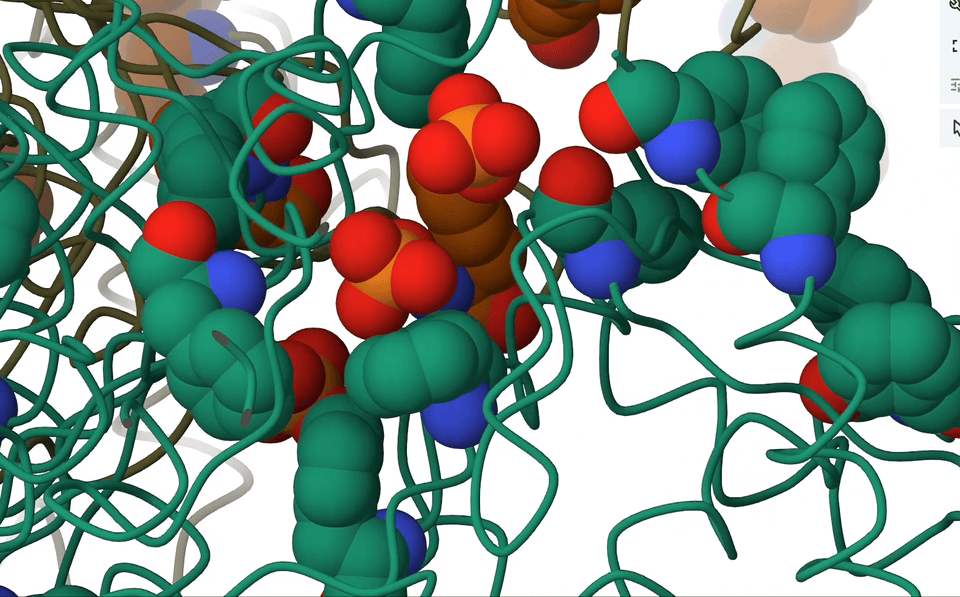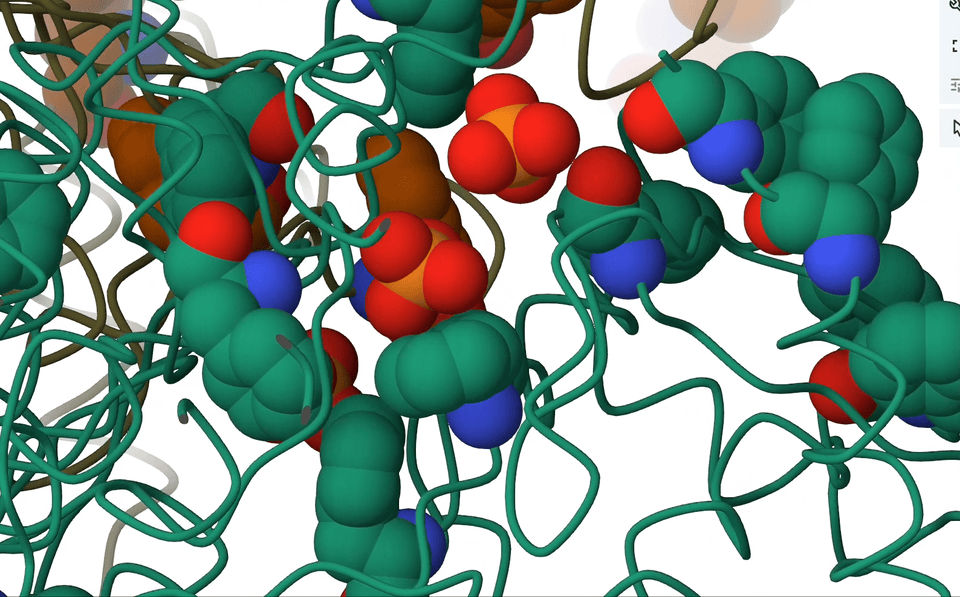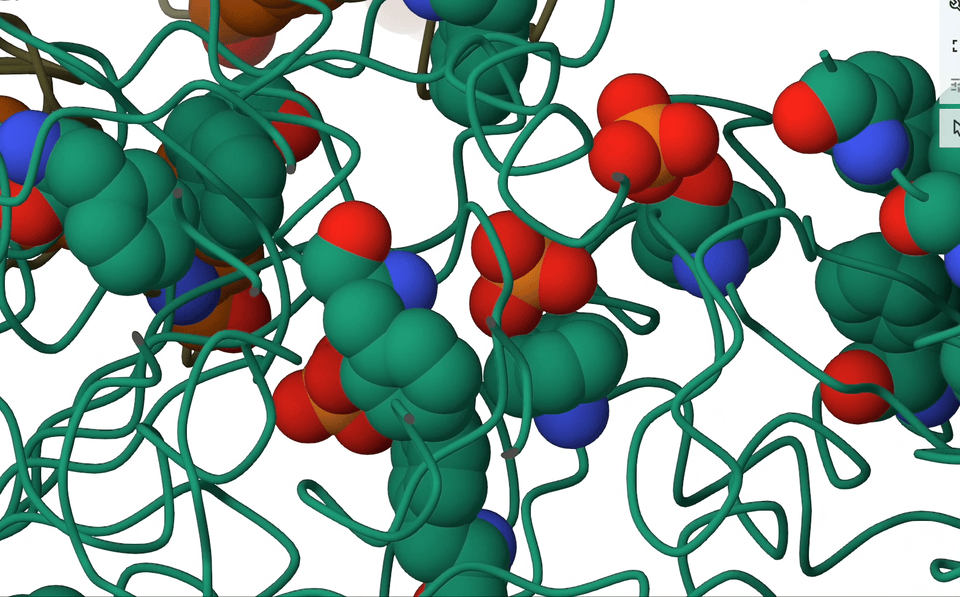
7. Anion-π interactions
Picture this: we have an anion, negatively charged and all, casually sitting atop aromatic rings that are electron-rich. Logically, one would assume they’d repel each other, right? But hold on, because in the right circumstances, they actually forge a delicate, attractive dance. It’s all about the dynamics of electrons, you see.
Ah, these little counter-intuitive wonders hold a special place in my heart! And I’m happy that Adhav and Saikrishnan referenced my 2011 study where I first proposed the existence of this unconventional noncovalent interaction in proteins.
Back in the day, I was knee-deep in exploring anion-π interactions within inorganic compounds and supramolecular chemistry. Quite established realms, I must say. Yet, that got me pondering: could this unique bonding possibly have a significant role to play in biological molecules?
A pioneering systematic search through the Protein Data Bank (PDB) showed that anion–π close contacts exist in experimental protein structures between the standard aromatic residues (Trp, Phe, Tyr, and His) and anions, such as chloride and phosphate.
Quinonero 2016
Think about it—proteins are teeming with aromatic rings and anions. So, with anticipation I rallied my brilliant colleagues, secured research funding, and embarked on an in-depth analysis of the entire PDB database. And guess what we found? Anion-π contacts in experimental protein structures! Although relatively weak, I concluded that these interactions could indeed have a role to play in biological processes.
Fast forward a few years, Quinonero & Co hailed my research as pioneering (oh that was a sweet discovery!) and confirmed my findings, showing that nearly 61% of protein structures feature at least one of these cute anion–π interactions.
Now, a decade since my initial paper, the field is buzzing with excitement as these interactions have firmly established themselves as essential elements of biopolymer structure and function.
More on From Atoms To Words:
▸ 60 Years in the Making: AlphaFold’s Historical Breakthrough in Protein Structure Prediction
▸ Large Language Models for Chemistry: Is the Beginning of a New Era?
▸ Water’s Hydrogen Bonds: What Makes Them Vital for Life As We Know It?
Noncovalent interactions in proteins to boost Structure Prediction
In our timeless pursuit of unraveling the enigmatic nature of protein function and properties based solely on sequence information, we have embarked on an arduous journey. Our desire to grasp the inner workings of enzymatic reactions, ligand binding, and protein folding has driven us forward. However, progress in structure-property prediction has been frustratingly sluggish.
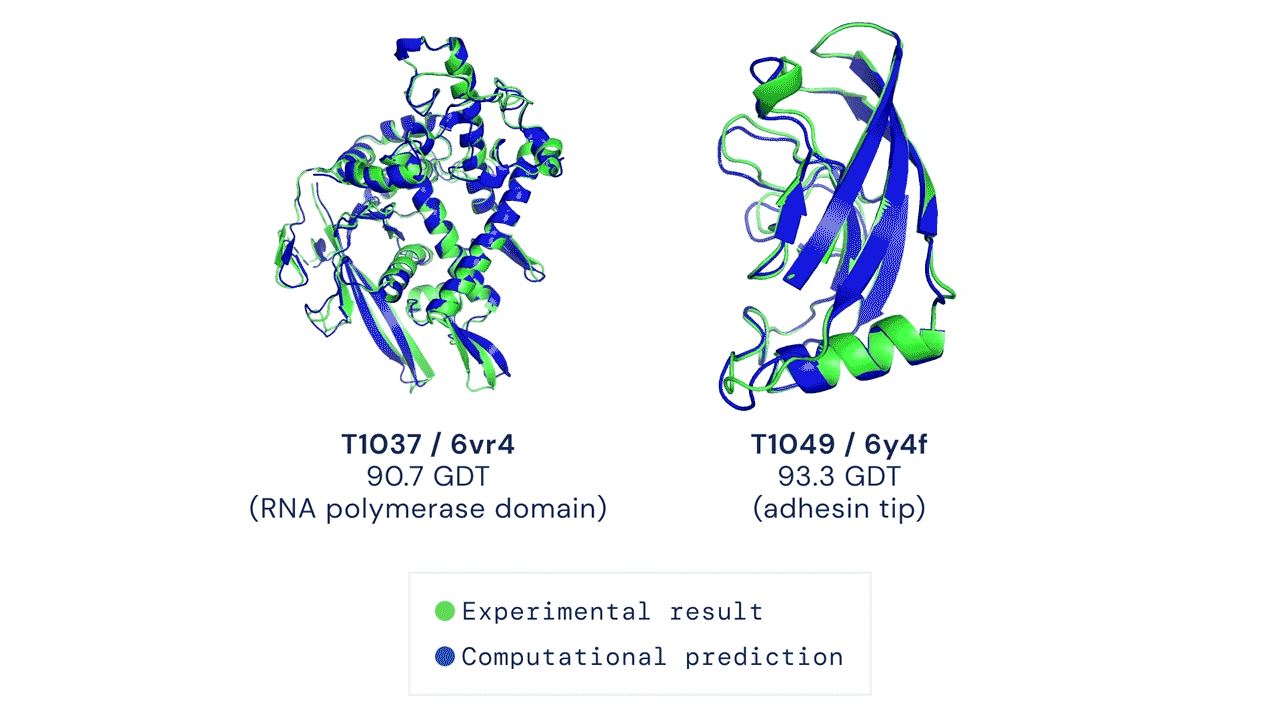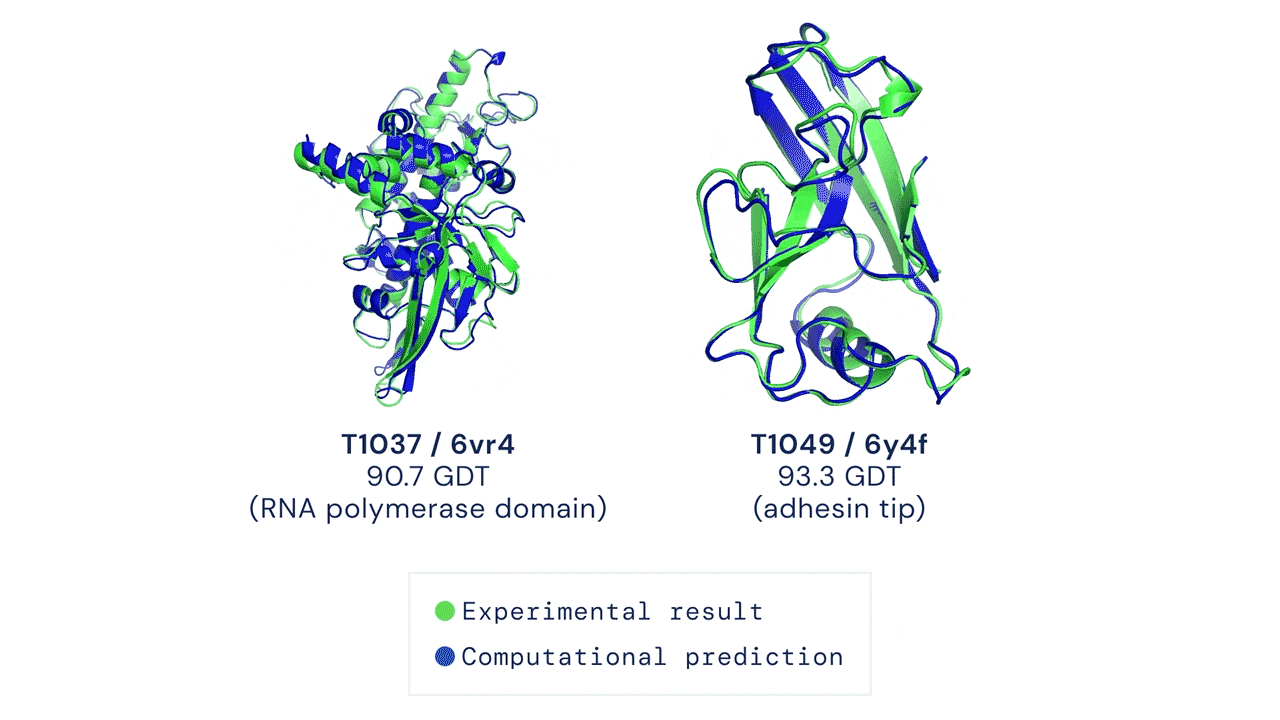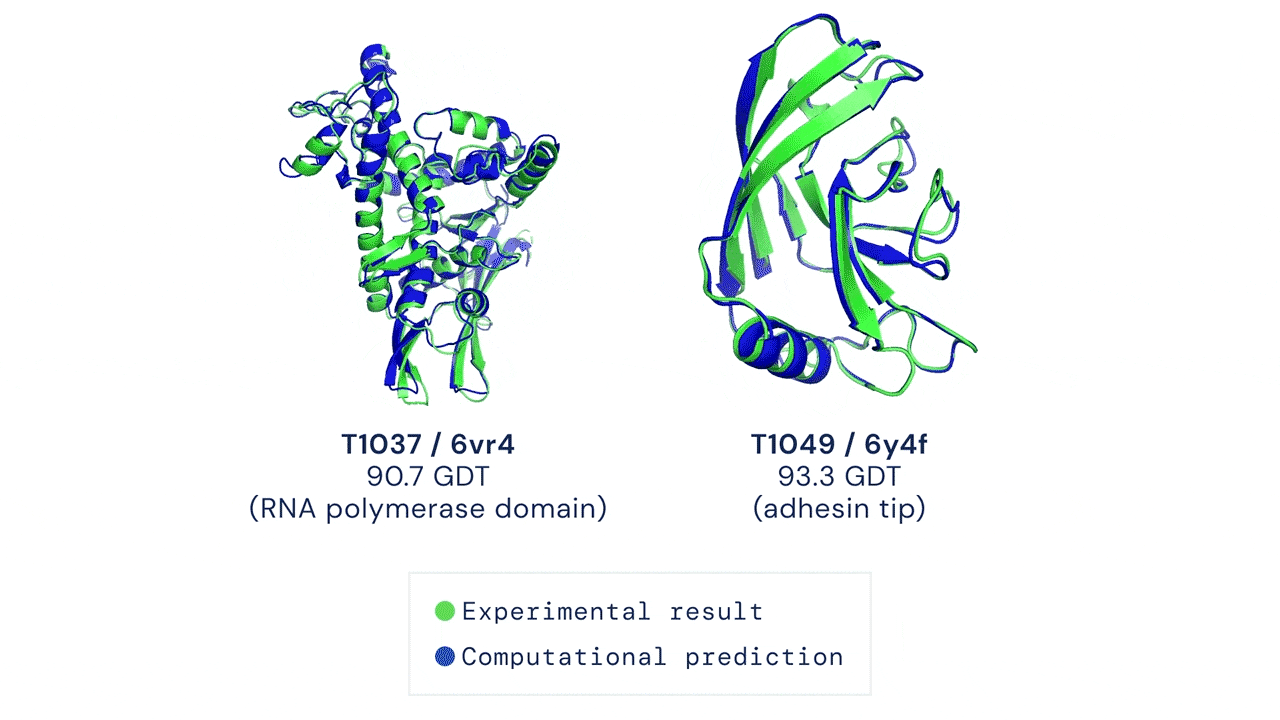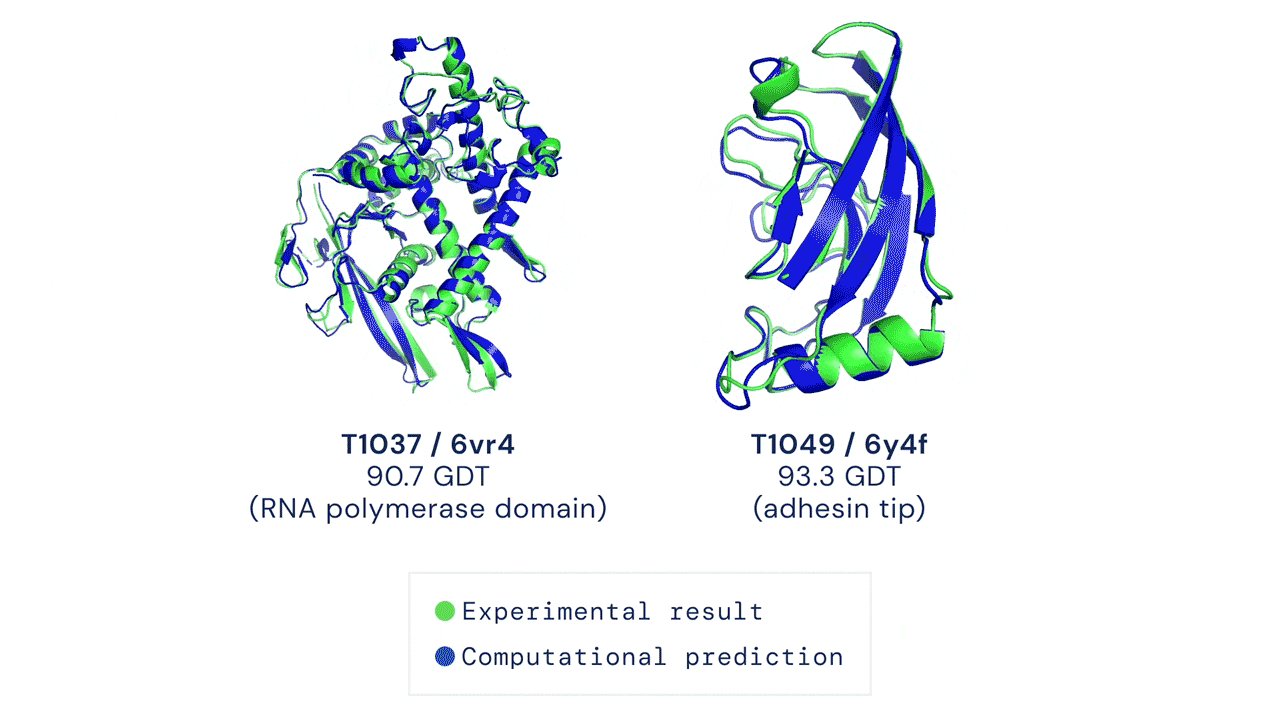
Yet, amidst the darkness, a glimmer of hope has emerged—machine learning. Welcome AlphaFold & Co, the trailblazers who have forever transformed protein structure prediction. These extraordinary AI models seamlessly integrate machine learning techniques with an expansive reservoir of sequence and structural data, empowering researchers to effortlessly obtain atomistic structures of proteins with unwavering confidence.
However, even this magnificent marvel of machine learning possesses its own inherent limitations. Its struggles lie in deciphering the intricacies of disordered proteins or complexes with ligands, cofactors, and nucleic acids.
Yet, do not be disheartened, for a promising path lies before us—a convergence of machine learning’s predictive prowess with molecular dynamics and quantum chemistry.
Now, to truly unlock this potential, we must diligently scrutinize the multitude of noncovalent interactions in proteins. Only by fully comprehending these forces at the atomistic level and appreciating the synergistic interplay between them, we can achieve our ambitious goal of decoding the protein function and properties from their sequence.
A Final personal touch
That article on anion-π interactions represents the culmination of my academic research. After that, I ventured into the world as an editor and later as a globetrotting sales & marketing manager for a medical device company. Though I found happiness in those roles, my gaze often wandered back to my golden years of anion-π research. From a distance, I convinced myself that I had left behind simulations, quantum chemistry, and proteins for good.
But, as it turns out, I was mistaken.
Nearly a year ago, I joined the team at Quantistry, and I found myself immersed once again in the world of simulations, engaging in research projects alongside industrial partners, and exploring the delights of all things “quantum.” I can honestly say that I’ve never found a more fulfilling job. Yet, deep within me, I continue to harbor a dream—a dream that someday, someone, perhaps inspired by that very article on anion-π interactions, will usher in a genuine scientific breakthrough.
Yes, I am a dreamer, but I’m not the only one.
If you enjoyed this dive into noncovalent Interactions in proteins and their marvelous features, I’d love to hear your thoughts. Agree, disagree, or have a totally wild theory of your own? Let’s connect! Subscribe to my LinkedIn newsletter and let’s keep the conversation rolling.